Dr. Ioan-Valeriu Grossu ioan.grossu@brahms.fizica.unibuc.ro
Medical Module Coordinator: Conf. Dr. Nicolae Verga: nicolae.verga@umfcd.ro
WPF "3D PictureBox" User Control by Mihai Opritescu: Research Gate
Graphic Design: Dr. Salma-Amalia El-Shamali Facebook Page
ABSTRACT
Visual tool for estimating the (fuzzy) fractal dimension of RGB images, medical three-channel images (CT, MRI), and N-Dimensional objects.
3D image reconstruction of data stored in csv file series, commeing from various sources (CT, MRI etc.).
INSTALLATION
OS: Windows 64-bit
Prerequisites: Microsoft .NET Framework 4.7.2
XCopy deployment strategy (simply copy/paste the application folder)
RELEASE NOTES
Hyper-Fractal Analysis v05
Improvements:
The application was rewritten in C# .Net in agreement with SOLID principles
The new version allows setting the fuzzy window filter boundaries
A new tool for identifying iso-fractal areas was implemented
The ND box-counting algorithm is not limited any more to 4 dimensions
Bug Fixes:
In the previous version (VB6), subunitary values were ignored in the fuzzy Box-Counting solution
Hyper-Fractal Analysis v06
Medical Module:
Possibility to open csv files containing three-channel images (see the attached examples from the App\Data folder)
New functionalities for facilitating slices comparison (e.g. CT native-contrast) by representing images on different RGB channels
Multi-channel generalized band-pass filter for csv three-channels images
New features:
Local [Fuzzy] Fractal Dimension export to csv
Bug Fixes:
Fuzzy Negative Band-Pass filter bug correction
Hyper-Fractal Analysis v07
3D Image Reconstruction:
The Platform Target was changet to x64 in order to avoid Out of Memory exceptions
Possibility to open csv file series containing three-channel images
Implementation of a "3D PictureBox" WPF user control for 3D image reconstruction
Hyper-Fractal Analysis v08
Fuzzy ND fractal analysis:
The current version is supporting fuzzy ND objects stored in specific csv files (.nfuz.csv extension).
The number of dimensions is limited only by the available computational resources
Various ND test scenarios for both classical and fuzzy implementations are covered by automatic tests
The user can display any (Xi, Xj) bidimensional projection of the ND object
The application was tested on 116D algal images obtained from the HYPSO-1 satellite
MAIN FUNCTIONALITIES
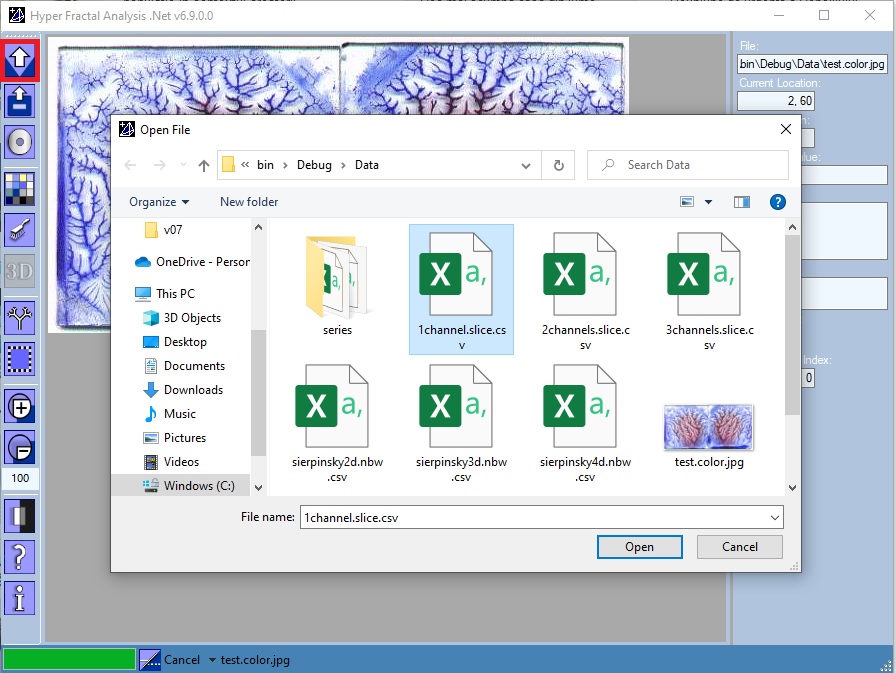
Open file
One could choose:
- An RGB image file (256 shades of gray is recommended)
- A "serial image" file: a comma separated values file with extension .nbw.csv, containing only the N-Dimensional coordinates of a set of "black" points
- A "serial image" fuzzy file: a comma separated values file with extension .nfuz.csv, containing the N-Dimensional coordinates together with the fuzzy degree of membership (a set of "gray" points)
- A three-channel csv file: a comma separated values file with extension .slice.csv, containing the matrices of up to 3 slices. Most commonly, we used the three channels for storing: native, arterial-contrast, and venous-contrast CT images.
|
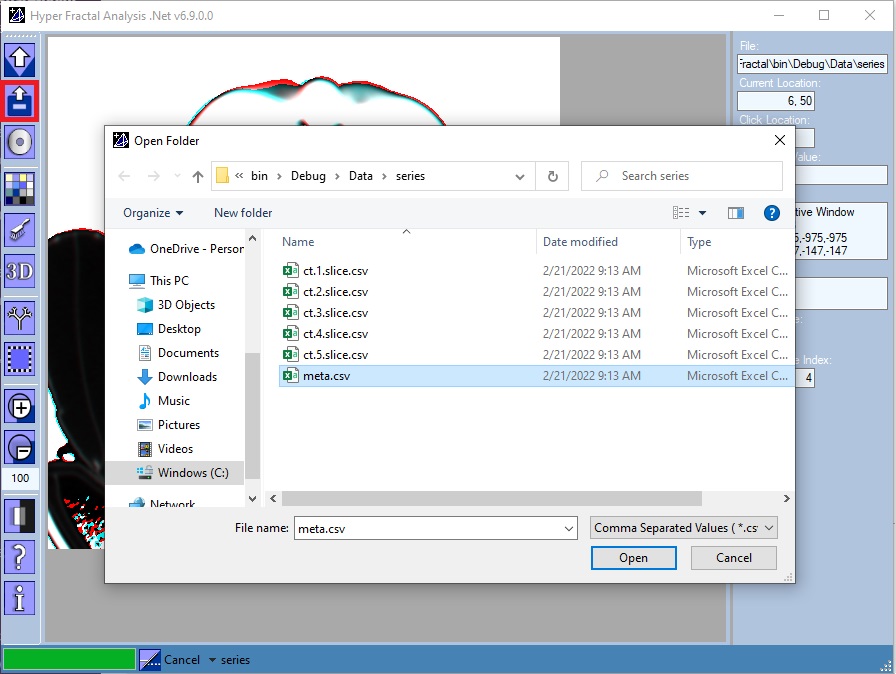
Open folder
One could choose:
- A folder containing a csv file series (a set of three-channel slices together with a csv metadata file)
|
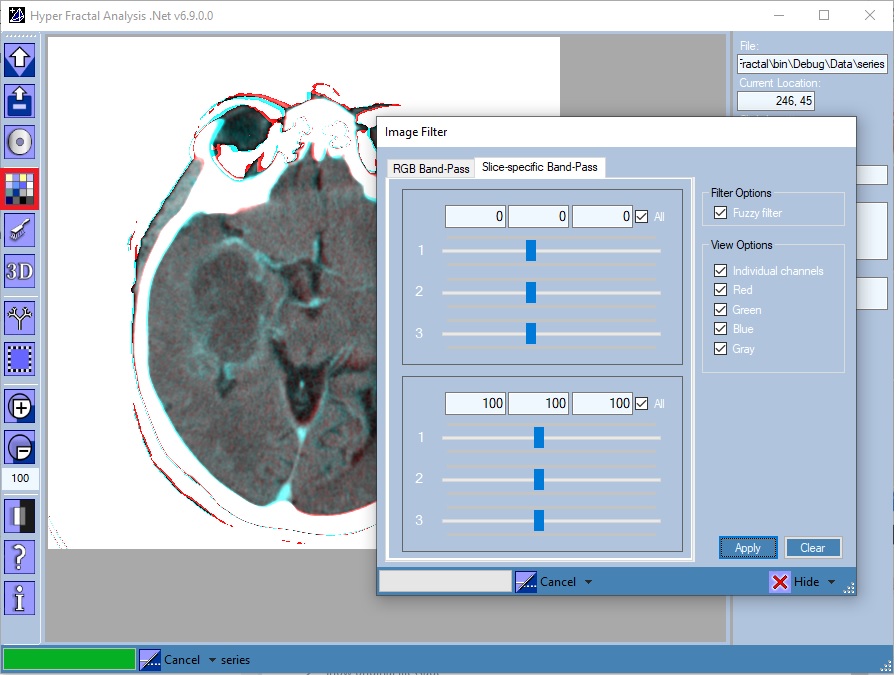
Image Filter
Open/Close the Image Filter tool
One could choose:
- A window filter: select only pixels in a specified window (band-pass filter)
- A fuzzy window filter: consider only the pixels in a specified window assigning also a weight between 0 (lower window level) and 1 (upper window level)
- Display any (Xi, Xj) bidimensional projection of the ND object
|
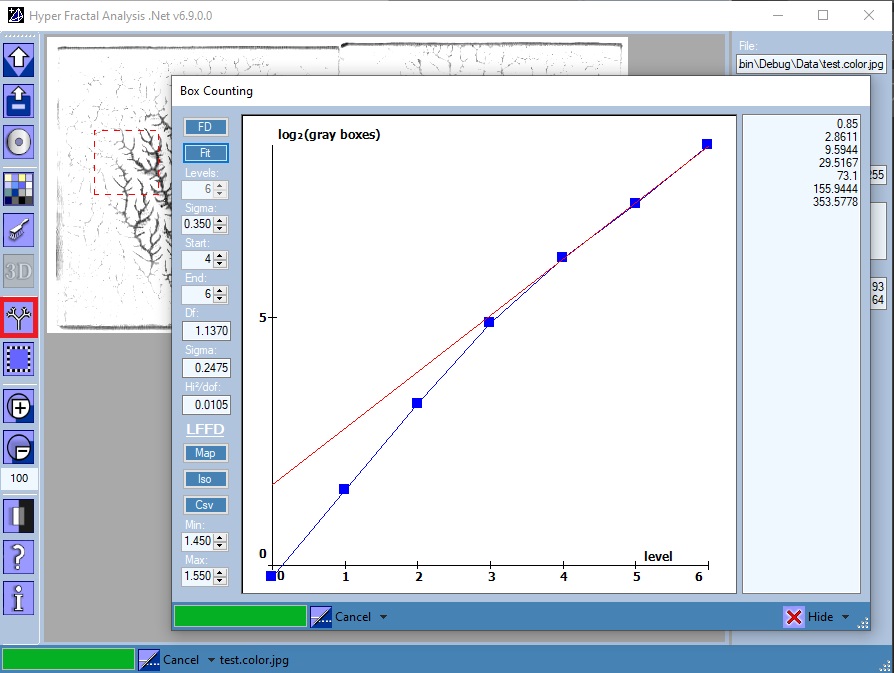
Box Counting
Open/Close the Box Counting tool
Features:
- FD Button
- Box-Counting algorithm for both 2D and ND images
- Parameters: the pixel error, and divisions level (available only for ND)
- Fit Button
- Linear fit for the previous Box-Counting solution
- Parameters: the solution range to be considered for the fit
- Map Button
- Draws a 256 shades of gray map of local fractal dimension values
- Parameters: the local fractal dimension range
- Iso Button
- Draws a black-wite map of local fractal dimension values
- Parameters: the local fractal dimension range
- This tool could be used for identifying the iso-fractal areas by choosing an enough small range
- Csv Button
- Exports the local [fuzzy] fractal dimension map to csv
|
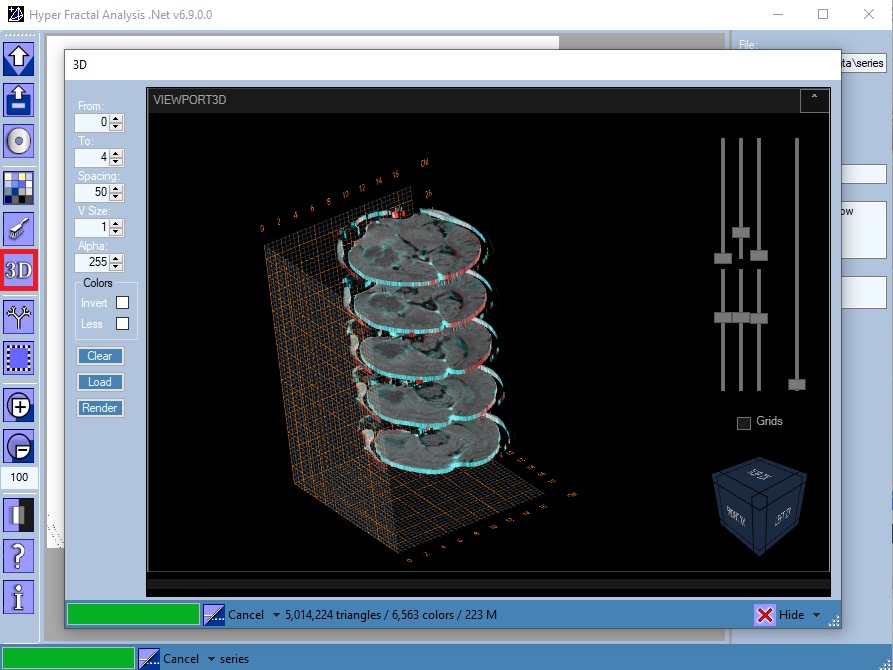
3D Reconstruction
Open/Close the 3D Reconstruction tool, available only for file series (see Open Folder). A filter is also required.
Parameters:
- The slices range (From - To zero-based indexes): used together with a rectangular image selection will define a 3D selection.
- The slices spacing: could be considered an an alternative to transparency
- The voxel's edge in pixels: edge > 1 => lower resolution (useful for testing)
- The voxel's transparency: 255 => no transparency (alpha compositing)
- Invert colors option
- Use less RGB colors option: rendering with many RGB colors could significantly affect performance
- The Load button is used for loading the voxels. Thus, in function of the calculated numbers of triangles and colors, the user could choose to Render or not the generated set.
|